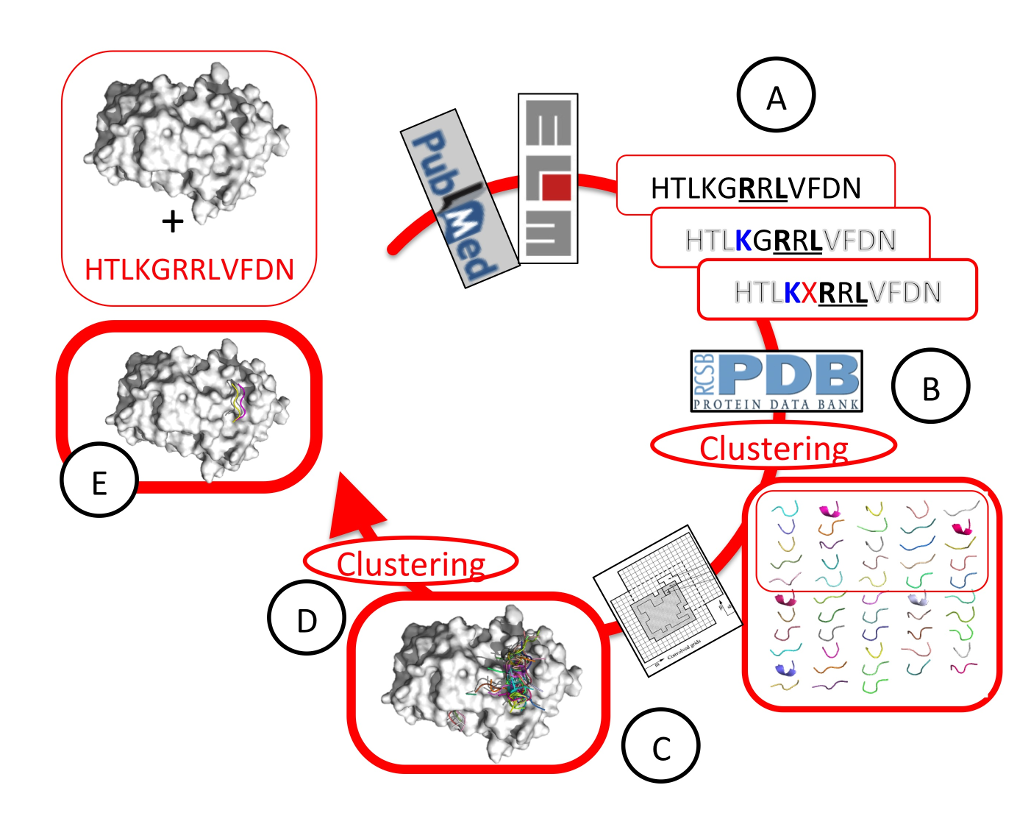
Protein-protein interactions play critical roles in cellular regulation and homeostasis. The accumulation of information on proteins, their modifications and their interactions in the present era of “big data” large-scale experiments provides an ever-increasing wealth of data that needs to be validated and understood in order to provide a unified framework of regulation in the cell. Around 40% or more of the crucial regulatory protein interactions are mediated by peptides, namely by the binding of a short, linear stretch located often in an unstructured region, to a globular domain in the partner protein . This estimate is even increased if we consider the many domain-domain interactions that are also mainly mediated by one short, “hot segment”.
At the level of an individual protein and its interactions, the atomic structure can teach us much about main underlying determinants, as well as provide a good starting point for the design of specific modulators of a given interaction . However, while the structural coverage of individual proteins has considerably increased lately, the percentage of solved structures of protein complexes remains low Docking protocols can bridge the gap between the number of solved structures of monomers and complexes.